Computational Modeling & Theory
Perisomatic biophysical single neuron models
Cells in the mammalian neocortex are diverse, and reproducing that diversity with computational models can be challenging. The experimental data collected in the Allen Cell Types Database provide valuable information for understanding the components of the mammalian neocortex. To continue toward that goal, hundreds of biophysically detailed models of single neurons recorded in vitro have also been created. These models were constructed using electrophysiological and morphological data collected from individual neurons.
The models characterize the firing properties of different cell types in the mouse visual cortex and human cortex, and relate them to generalized biophysical mechanisms governing this firing. Biophysically detailed models may also generalize to different types of input more naturally than more abstract models. The models presented in the Allen Cell Types Database differ from other collections of models in that they are built from data obtained in a highly standardized way and are presented alongside the original experimental data upon which they are based.
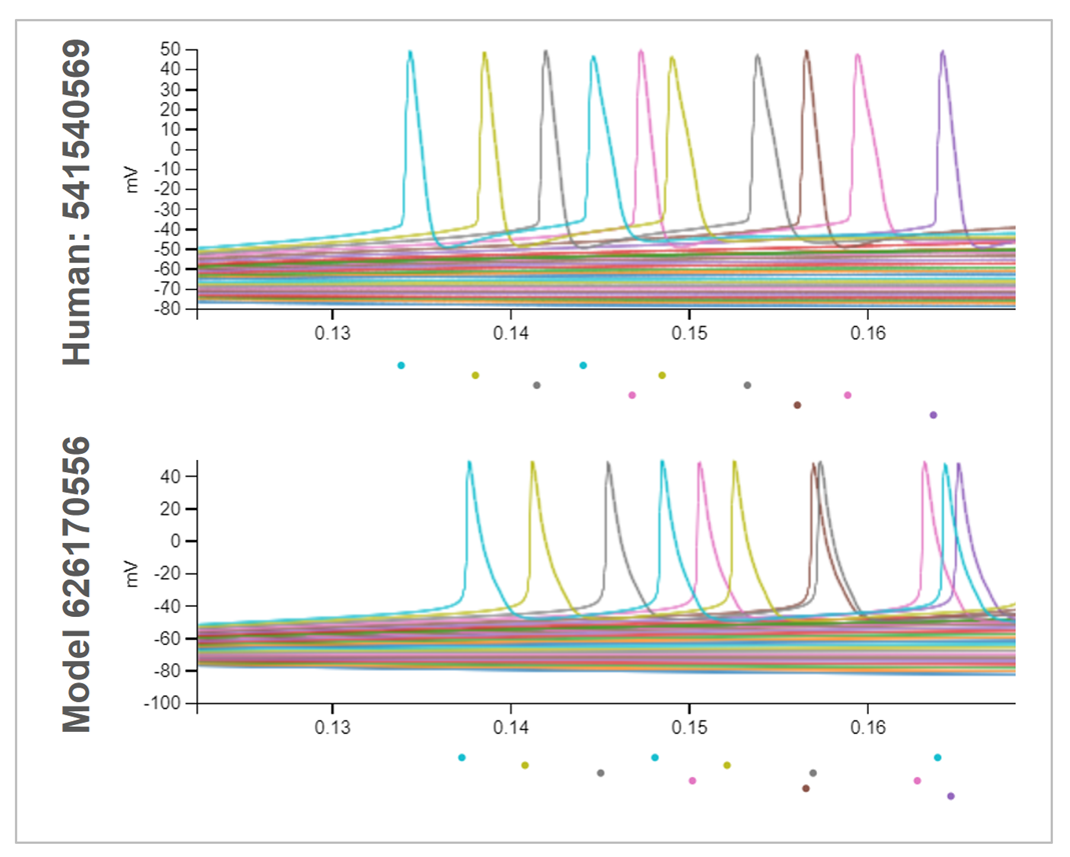
The biophysically detailed models were fit with multiple channels placed at the cell soma, using data from individual neurons in the Allen Cell Types Database. Measurements from reconstructed cell morphology are used, ionic conductances are placed at the soma, and the conductance densities are optimized using a genetic algorithm to train the models to the experimentally-measured responses to step currents of a single amplitude. The model is tested in NEURON by applying all the other stimuli applied to the experimental cell being modeled.
Access data with perisomatic models
The models are viewable alongside experimental data in the Allen Cell Types Database. Select data with models by filtering for 'Perisomatic Biophysical Model,' and access data and models for individual cells in the Cell Results panels. The Electrophysiology Summary displays experimental and modeled data, which can also be downloaded. Code to run the models is provided through the AllenSDK. The optimization code is available on GitHub.
Your browser is out-of-date!
Update your browser to view this website correctly.